AI is reshaping life science research, but poor reporting of AI and ML methodologies undermines transparency and reproducibility. The ELIXIR Machine Learning Focus Group has developed the DOME Recommendations and the DOME Registry to bring clarity, standardization and accountability to AI/ML research. Read on to discover how DOME is helping scientists, reviewers and publishers understand what’s on the ‘ingredients list’ and build trust in AI/ML methods.

Innovation without ingredients
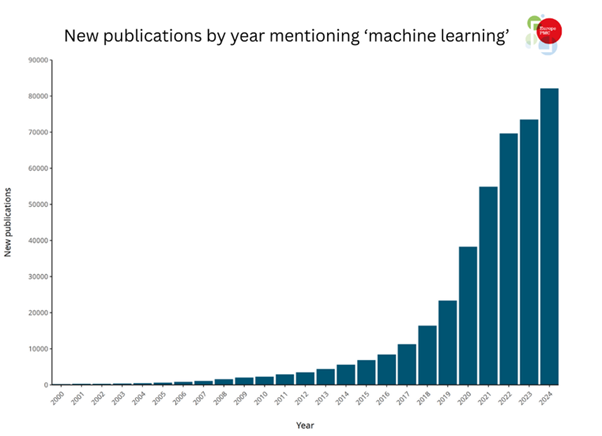
Artificial intelligence (AI) and machine learning (ML) methods are transforming scientific discovery. Accelerated by access to high-power graphics process units, FAIR (findable, accessible, interoperable and reusable) data and success stories such as AlphaFold 2,1,2 more research articles using AI/ML methodologies are being published than ever before (Figure 1).
However, it is extremely challenging to adequately describe complex AI/ML methodologies in traditional publication formats, and there is limited standardization in reporting requirements across publishers. As a result, many AI/ML papers lack sufficient methodological detail, making it difficult for reviewers and readers to understand, reproduce or build on the work presented.4
Consider how we make decisions about food. Many pre-packaged foods have nutrition labels that provide key information about ingredients, serving size, nutrients present and daily recommended values. We can use this information to compare products and make informed choices about the groceries we buy.
However, there is no such ‘nutrition label’ for AI/ML methods reporting, and the absence of clear guidance on what should be disclosed to ensure robust reporting has resulted in a field plagued by poorly described publications that lack information on datasets, provenance, data leakage between testing and training sets, model software availability and unfair evaluations.
The chefs
This is where ELIXIR Europe stepped in. ELIXIR is an intergovernmental organization that brings together life science resources – including databases, software tools and supercomputers – to form a single, Europe-wide research infrastructure. Within the ELIXIR community are a series of focus groups designed to bring together interested parties from across the organization to discuss emerging areas of interest and develop action strategies.
Between October 2019 and September 2025, the ELIXIR Machine Learning Focus Group came together to develop ML standards, support reproducibility and benchmark ML tools. Part of this work included the development of the DOME Recommendations for reporting supervised ML-based analysis of biological studies.5
The DOME Recommendations
The DOME Recommendations are much like a nutrition label – but for ML research methodologies. Just as a nutrition label breaks down the composition of food into clear, standardized nutritional categories, the DOME Recommendations break down the essential components of ML methodologies into four transparent pillars: Data, Optimization, Model and Evaluation (below). Instead of calories, fats and sugars, the DOME Recommendations include 20 items researchers should report to ensure their research is transparent and reproducible, including how data were curated and split, how models were tuned, which architectures were used and how performance was assessed.
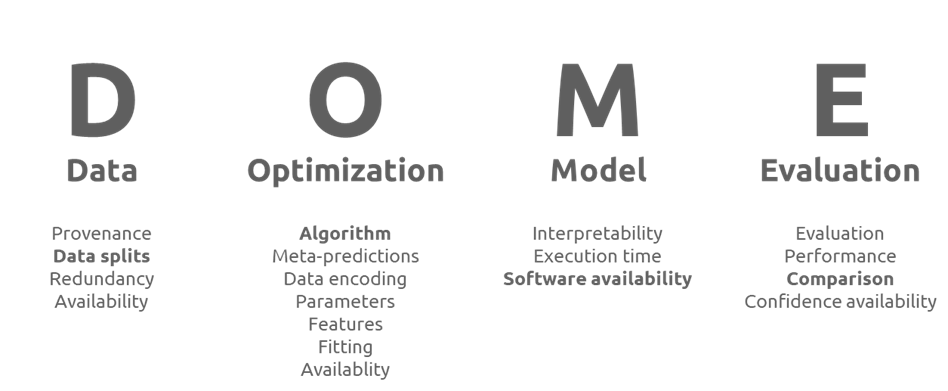

The structured, question-based framework breaks down the ‘ingredients’ of the AI/ML methods and helps researchers to report them in a way that makes it easy for readers to see what went into the model, how it was prepared and whether the reporting meets basic quality expectations. Much like comparing two cereal boxes for fibre content or added sugars, researchers and readers can compare two ML papers using their DOME‑related information to assess transparency, reproducibility and overall methodological robustness.
The DOME Registry
If the DOME Recommendations are the ingredients list, the DOME Registry6 is the recipe book and food critic rolled into one. The DOME Registry is used to curate and annotate submitted AI/ML methodologies according to how transparently they report the essential details identified by the DOME Recommendations. After curation, each methodology is assigned an overall compliance score that allows researchers, peer reviewers and editors to quickly assess whether they adhere to best practice recommendations and comply with editorial policies based on the curation.
Following a successful pilot integration into the publishing workflows of GigaScience and GigaByte, the registry is open for adoption by more publishers. We actively encourage engagement to gather insights that will help expand adoption across life science domains and optimize the service for the needs of specific communities. We also support community and retrospective curation to ensure older, critical models are not left behind.
To understand exactly how the Registry works in practice, it is helpful to look at the different chefs in the kitchen.
- Authors – annotate and submit methods: during the submission of a manuscript, authors can register their AI/ML methodology through curation and annotation in the DOME Registry. This generates a unique identifier and acts as a compliance report, effectively acting as a ‘quality seal’ for their ingredients list, which they can include in their paper to demonstrate transparency and support reuse by others.
- Researchers – discover methods: for those looking to reuse or build upon existing work, the registry serves as a curated cookbook. Researchers can browse registered methods to find well-documented models, compare ‘recipes’ based on their compliance scores, and ensure they are selecting robust methods for their own experiments.
- Journals – mandate DOME use: publishers and editors can integrate the registry into their submission workflows. By adopting the DOME Registry as a publishing requirement, journals can help verify that the ‘nutritional information’ of a submission meets their standards, streamlining the review process and reducing the burden on peer reviewers who can best assess publication methods.
- DOME Registry staff – support the infrastructure: behind the scenes, the registry staff act as the head chefs and health inspectors. They maintain the infrastructure, validate community annotations, and ensure that the ‘food safety standards’ (the interoperability standards and reporting criteria) remain up to date with the rapidly evolving AI landscape.
The future of the DOME Registry
Looking ahead, we are set to soon release our DOME Registry roadmap guided by the service’s external Scientific Advisory Board. Our future roadmap includes coverage across areas such as:
- standardization: using ontologies and free-text reduction to make DOME’s data machine-readable and FAIR
- frontier standards: considering adoption and integration of emerging standards such as Croissant and FAIR4ML to ensure interoperability for cross-resource asset exchange
- discoverability: creating a bidirectional connection between research outputs archived in Europe PMC and methodologies stored in the DOME Registry, powered by the EBI search programme to enhance discoverability
- scaling: exploring alternative scaling approaches for method generation through informal pilot projects with the AI4EOSC team based at Consejo Superior de Investigaciones Científicas (CSIC), Spain.
We warmly welcome new engagement from industry partners and publishers. If you’re interested in learning more about DOME, including how to use it at your organization, contact the team at contact@dome-ml.org. To stay updated on our progress, follow the DOME news and events.
By adopting DOME, the life sciences community can ensure that innovations built on AI/ML methodologies remains transparent, reproducible and trustworthy, building a stronger foundation for future breakthroughs.
References
- Jumper J, Evans R, Pritzel A et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021;596:583–9. https://doi.org/10.1038/s41586-021-03819-2.
- Callaway E. Chemistry Nobel goes to developers of AlphaFold AI that predicts protein structures. Nature 2024;634:525–6. https://doi.org/10.1038/d41586-024-03214-7.
- Farrell G, Attafi OA, Psomopoulos F et al. DOME Registry supporting ML transparency and reproducibility in the life sciences – presentation (ISMB/ECCB 2025). 2025. Available from: https://doi.org/10.5281/zenodo.15807199 (Accessed: 30 January 2026).
- Ball P. Is AI leading to a reproducibility crisis in science? Nature 2023;624:22–5. https://doi.org/10.1038/d41586-023-03817-6.
- Walsh I, Fishman D, Garcia-Gasulla D et al. DOME: recommendations for supervised machine learning validation in biology. Nat Methods 2021;18:1122–7. https://doi.org/10.1038/s41592-021-01205-4.
- Attafi OA, Clementel D, Kyritsis K et al. DOME Registry: implementing community-wide recommendations for reporting supervised machine learning in biology. GigaScience 2024;13:giae094. https://doi.org/10.1093/gigascience/giae094.
The views expressed in this blog post are those of the author and do not necessarily reflect those of Open Pharma or its Members and Supporters.
Gavin Farrell is a PhD student at the University of Padova, Italy.



